Engineering CAR-T Cells to Target Intracellular Cancer Drivers
Speaker:
Mark Yarmarkovich, PhD
Assistant Professor, Dept. of Pathology
Perlmutter Cancer Center, NYU Grossman School of Medicine
Abstract:
Dr. Yarmarkovich’s research is centered around the development of personalized immuno- therapies (CAR T cells, TCRs and BiTEs) for all cancer patients. His groups work spans the entire workflow from target discovery to clinical application. This research situated at the multidisciplinary intersection of genomics, proteomics, immunology, protein engineering and computational biology to discover novel tumor-specific targets and engineering the target-specific immune receptors. Dr. Yarmarkovich’s team has discovered novel tumor targets from previously undruggable proteins and has developed a novel class of "peptide-centric" (PC) CAR T cells that dramatically expand the immunotherapeutically targetable landscape and the addressable patient population. These CAR T constructs completely eradicate tumors in preclinical models and are entering clinical trials in this year. Furthermore, his lab is currently expanding these techniques to other cancers and developing new technologies for the development of personalized immunotherapies.
After receiving a BS degree in Biomedical Engineering from the University of Wisconsin – Madison in 2007, Dr. Yarmarkovich joined Genentech. There he worked in the antibody-drug conjugate (ADC) group to develop novel delivery systems for therapeutic agents used in oncology. In 2012 he started to pursed a PhD degree in Cell and Molecular Biology at the University of Pennsylvania. From 2019 to 2022 he was a postdoctoral fellow with John Maris at the Children's Hospital of Philadelphia. During that time, he co-founded two companies, Tantigen Bio and Hula Therapeutics, that focus on the development of new immunotherapies for cancer. 9 months ago, in January 2023, he joined NYU as an Assistant Professor of Pathology.
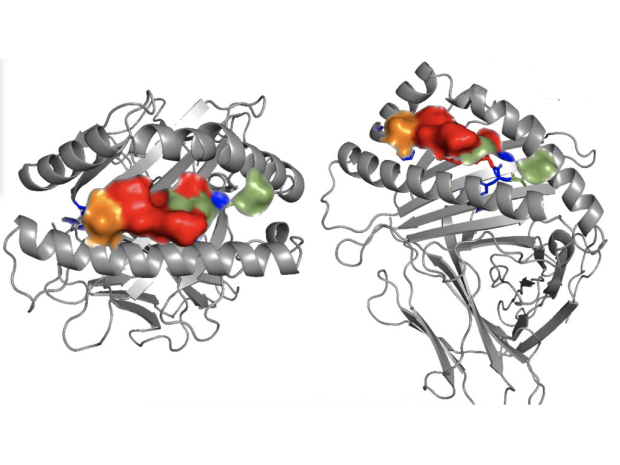
(a) PHOX2B–HLA-A*24:02 crystal structure and models of PHOX2B in complex with HLA-A*23:01, HLA-B*14:02 and HLA-C*07:02. (b) Charged and polar R151, Q155 and R69 residues of HLA-C*07:02 align with key 10LH inter-action residues I5, R6 and I7 (MHC residues in blue and PHOX2B–10LH interaction residues in red).

