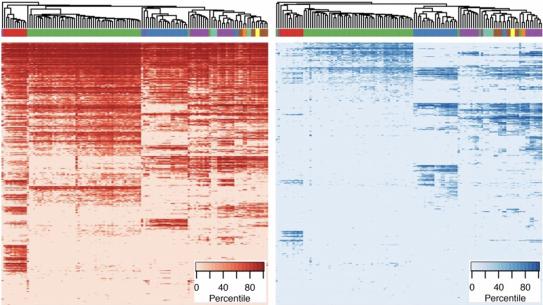
Epigenomic Maps of the Human Heart

Speaker:
Aravinda Chakravarti, PhD
Muriel G. and George W. Singer Professor of Neuroscience and Physiology
Professor, Department of Medicine & Institute for Systems Genetic
Director, Center for Human Genetics and Genomics
NYU Grossman School of Medicine
Abstract:
Cis-regulatory elements (CRE), short DNA sequences through which transcription factors (TFs) exert regulatory control on gene expression, are postulated to be the major sites of causal sequence variation underlying the genetics of complex traits and diseases. We present integrative analyses, combining high-throughput genomic and epigenomic data with sequence-based computations, to identify the causal transcriptional components in a given tissue. We use data on adult human hearts to demonstrate that (1) sequence-based predictions detect numerous, active, tissue-specific CREs missed by experimental observations, (2) learned sequence features identify the cognate TFs, (3) CRE variants are specifically associated with cardiac gene expression, and (4) a significant fraction of the heritability of exemplar cardiac traits (QT interval, blood pressure, pulse rate) is attributable to these variants. This general systems approach can thus identify candidate causal variants and the components of gene regulatory networks (GRN) to enable understanding of the mechanisms of complex disorders on a tissue- or cell-type basis.
Dr. Chakravarti has been a key participant in the Human Genome Project, the International HapMap Project and the 1000 Genomes Project. His research attempts to understand the molecular basis of multifactorial disease. He was awarded the 2013 William Allan Award by the American Society of Human Genetics and the 2018 Chen Award by the Human Genome Organization.